Chapter 3 Randomized Inference
Adapted from Cunningham (2021)
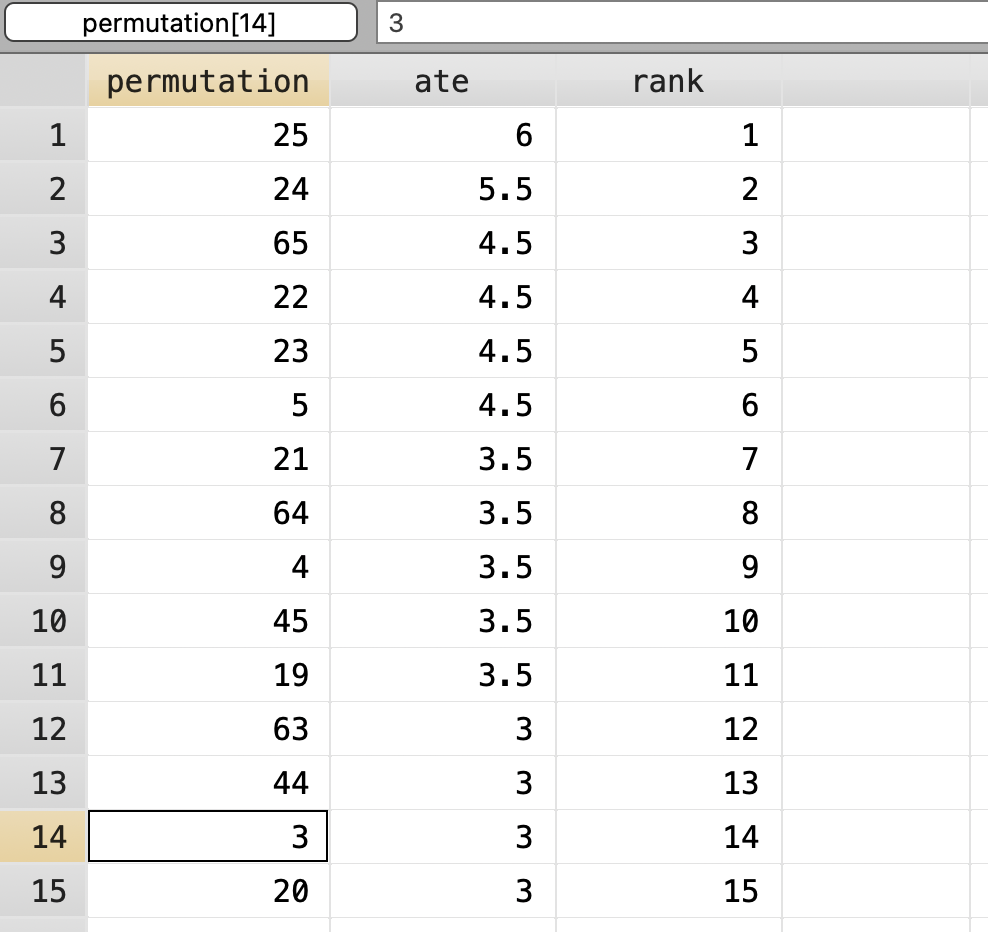
We will use a placebo-based test to calculate exact p-values. We need every single permutation of treatment to calculate the exact p-values from the ATE.
3.1 Set up the Dataset
First we will set the seed so we can replicate our randomized results
Next we will bring in our
cd "/Users/Sam/Desktop/Econ 672/Data"
use ri.dta, clear
tempfile ri
gen id = _n
save "`ri'", replace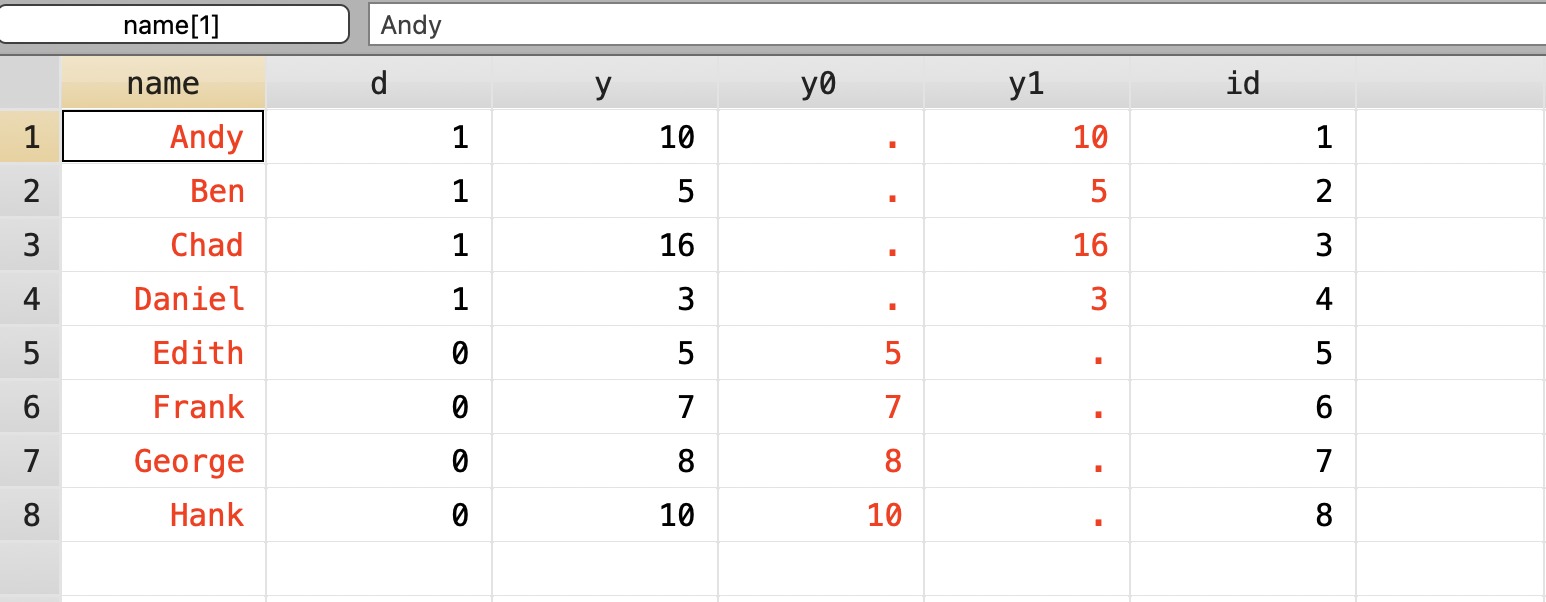 Next we will create all combinations from our eight observations. We will need to install the percom package into stata.
Next we will create all combinations from our eight observations. We will need to install the percom package into stata.
Out of 8 observations, we get 70 permutations. We can use the combin command to get these permutations. From the help combin, the combin command generates \(k\) unique combinations from a set of \(n\) objects, where \(k\) is not greater than \(5 (2 =< k <= 5)\). Program first arranges \(k\) possible permutations and then retains unique combinations. It supports both string and numeric variables.
combination: \(\frac{n!}{k!(n-k)!}\)
We will tell the combin command that we want 4 combinations from our 8 observations.
k = 4
N = 8
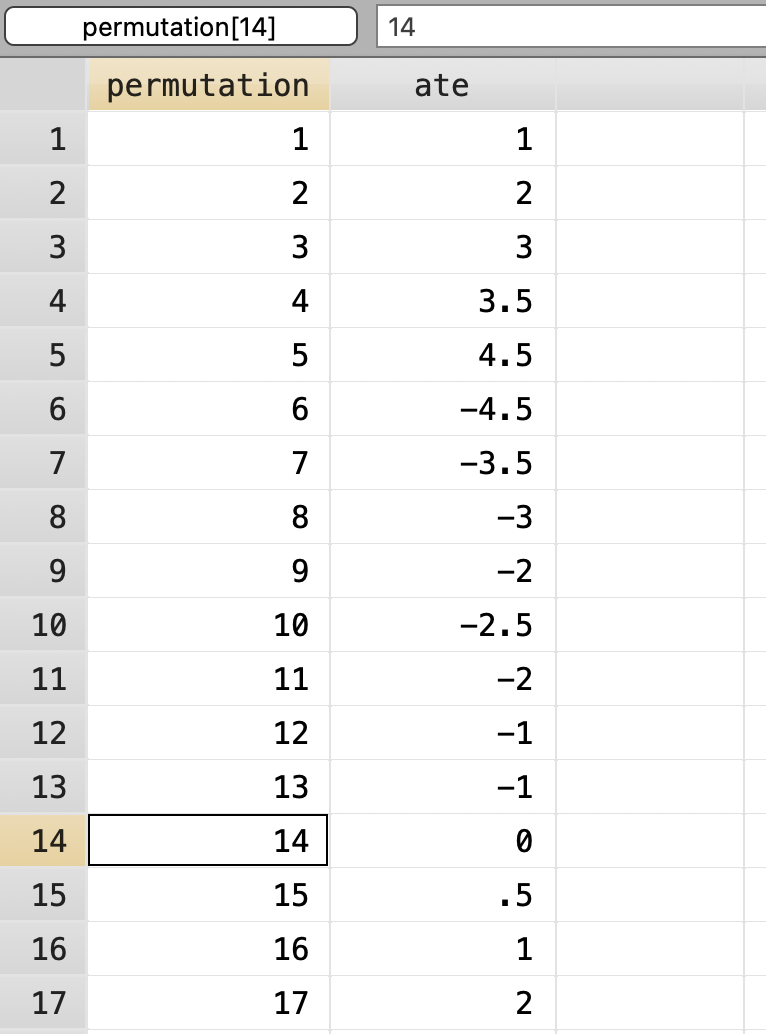
Combinations Formed: 70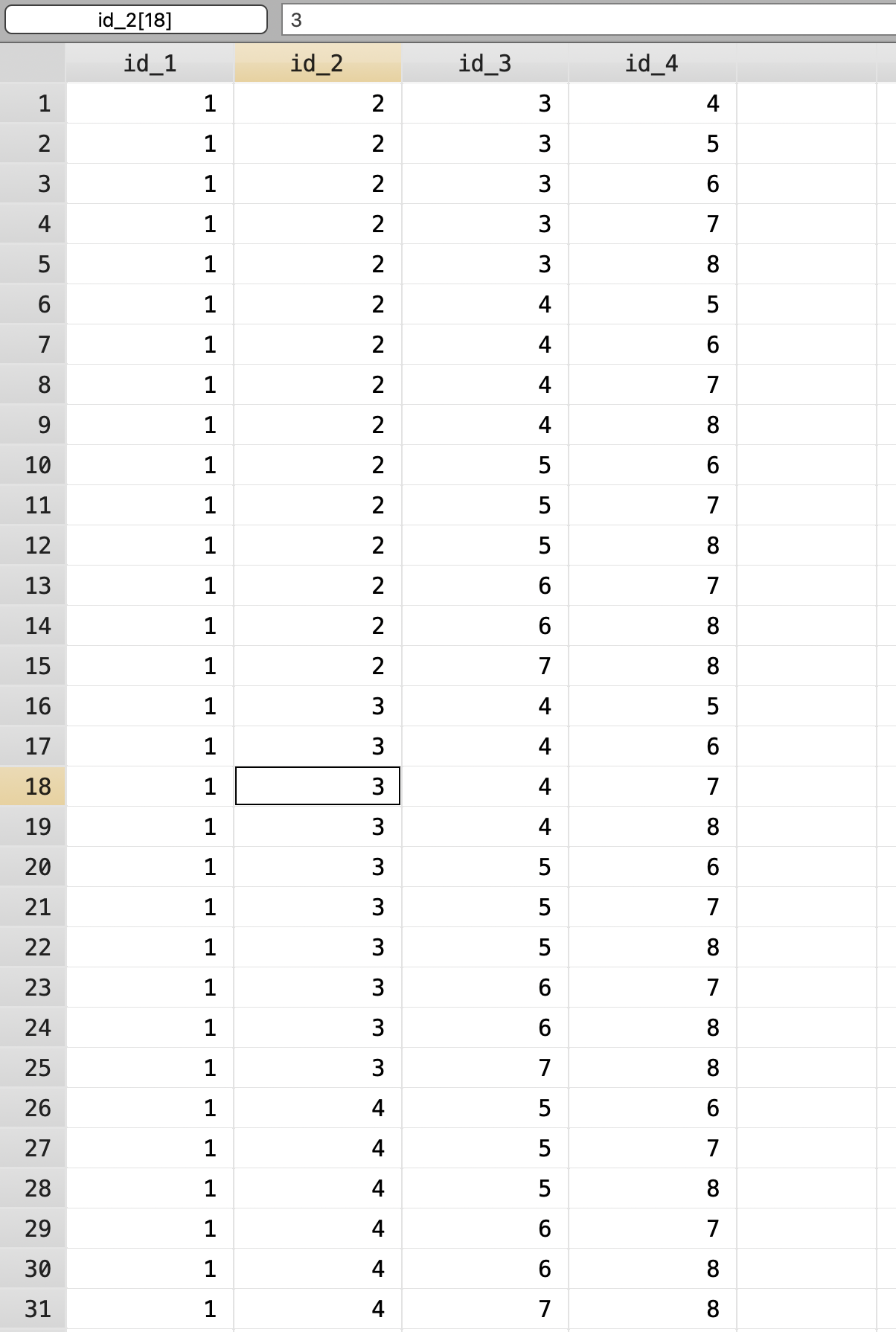 Next we will label the permutations 1 to 70.
Next we will label the permutations 1 to 70.
gen permutation = _n
tempfile combo
save "`combo'", replace
forvalue i =1/4 {
rename id_`i' treated`i'
}Next we will merge our permutations with our dataset of potential outcomes
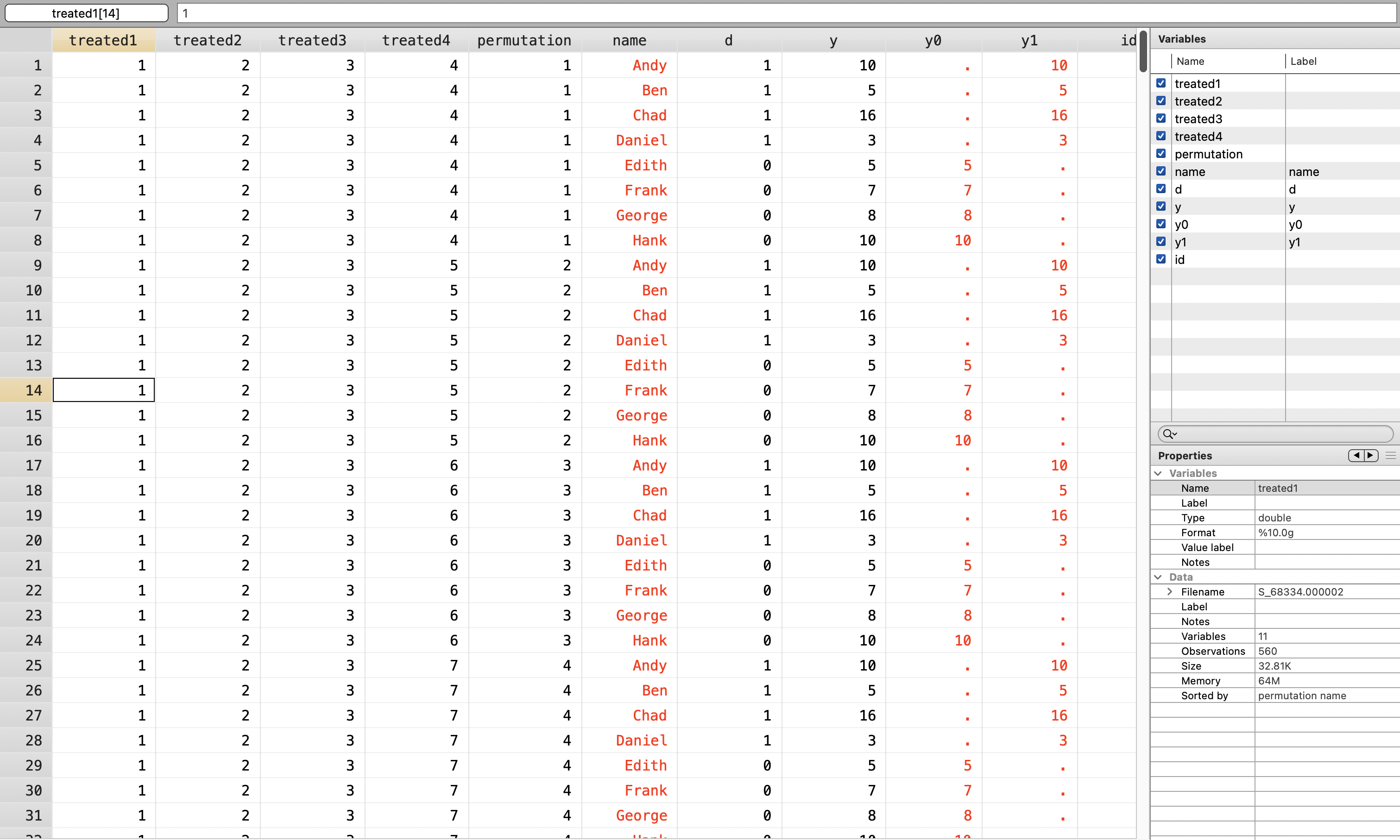
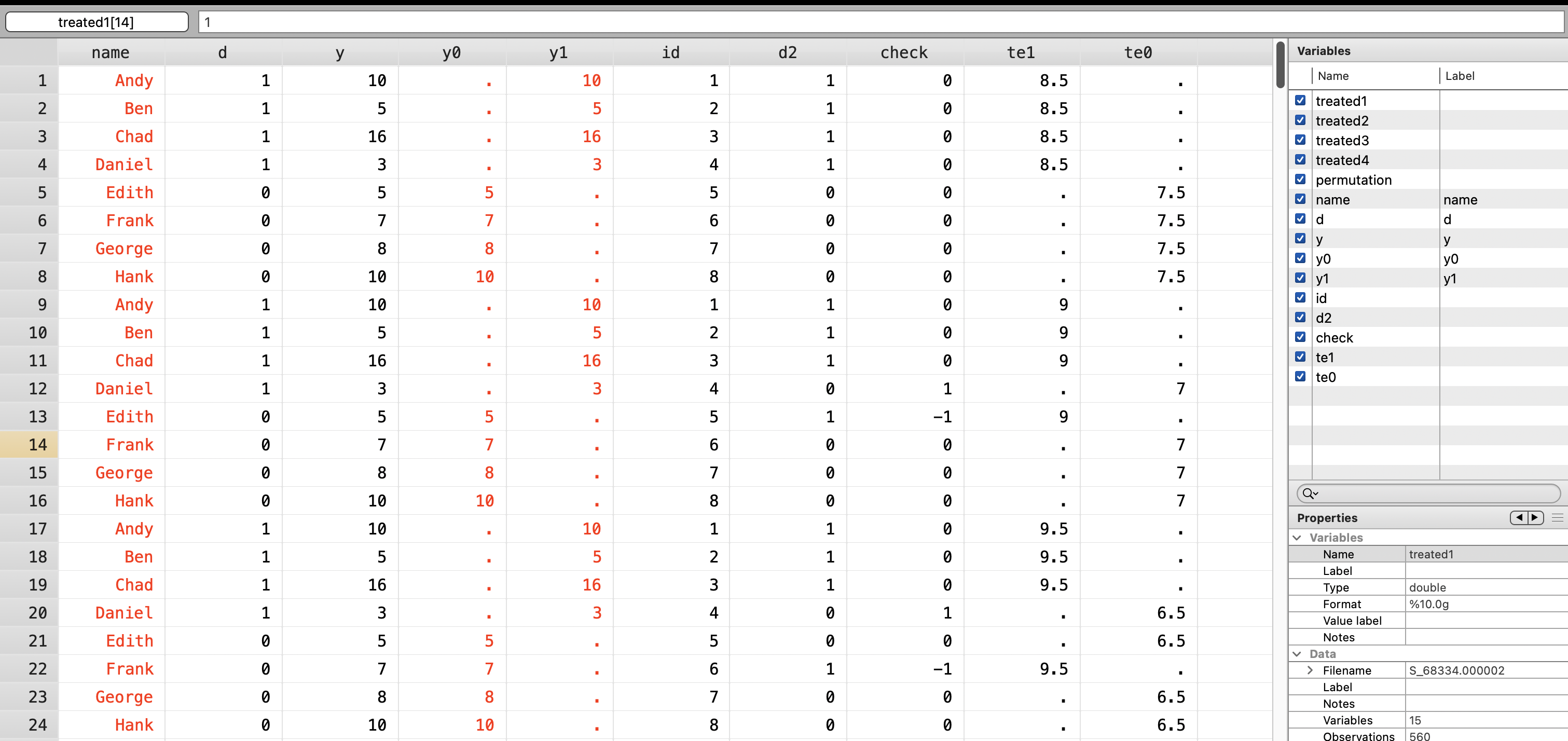
3.2 Randomize Assignment to Treatment
Next we will randomized treatment and calculate other test statistics.
First, we create a fake treatment variable \(d2\)
gen d2 = .
replace d2 = 1 if id == treated1 | id == treated2 | id == treated3 | id == treated4
replace d2 = 0 if ~(id == treated1 | id == treated2 | id == treated3 | id == treated4)
gen check = d - d2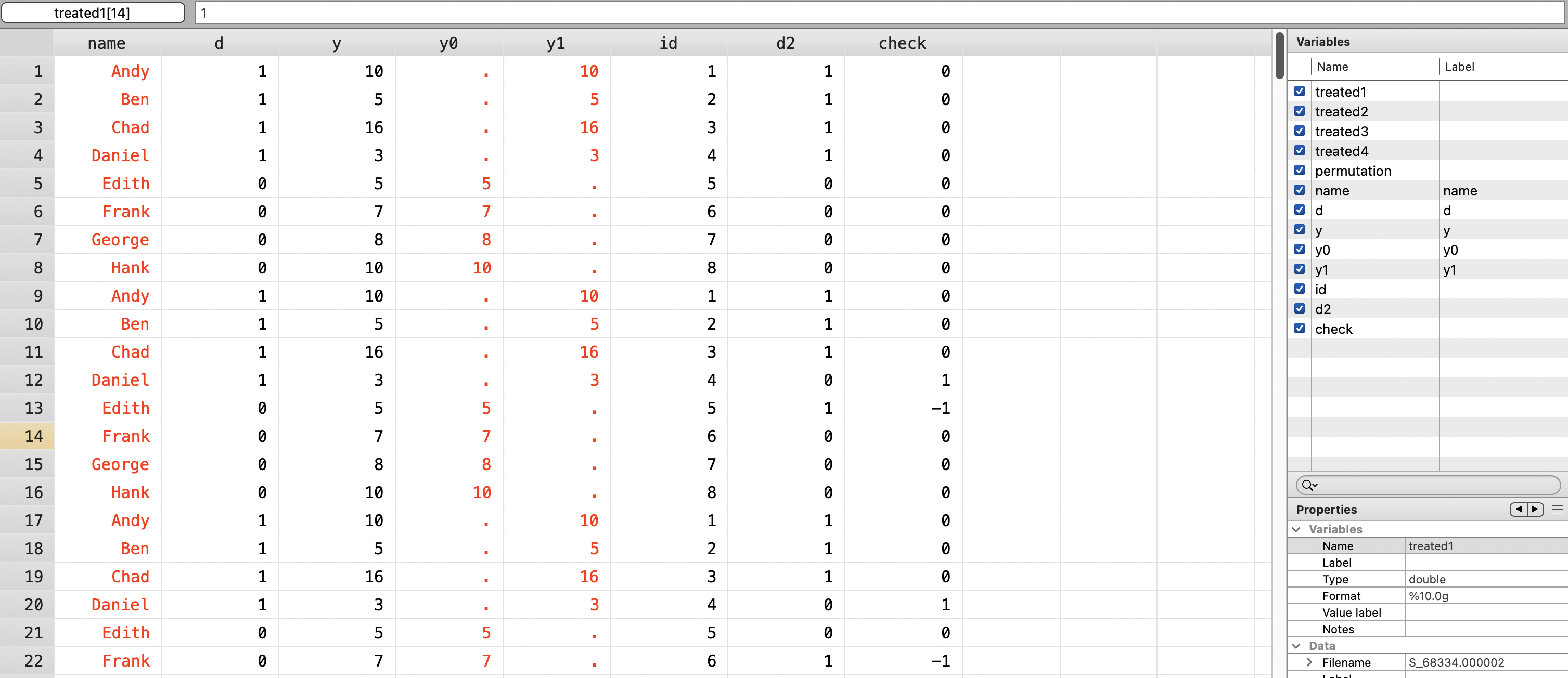
Next, we will calculate true effect using absolute value of simple difference in outcomes \((SDO)\).